BioByte 154: DefensePredictor Uncovers Bacterial Defense Mechanisms, MaxToki Reasons Across Cell Stage Trajectories to Predict Aging, and Engineered Trogocytosis as a Programmable Delivery Mechanism
Welcome to Decoding Bio’s BioByte: each week our writing collective highlight notable news—from the latest scientific papers to the latest funding rounds—and everything in between. All in one place.
What we read
Papers
DefensePredictor: A machine learning model to discover prokaryotic immune systems [Deweirdt et al., Science, April 2026]
Why it matters: DefensePredictor utilizes protein language model embeddings to uncover new bacterial defense systems which are overlooked with classic sequence-homology based methods. Deweirdt et al. uncover new families of anti-phage defense proteins that have the potential to be repurposed to powerful genetic engineering technologies, just like that of CRISPR-Cas9.
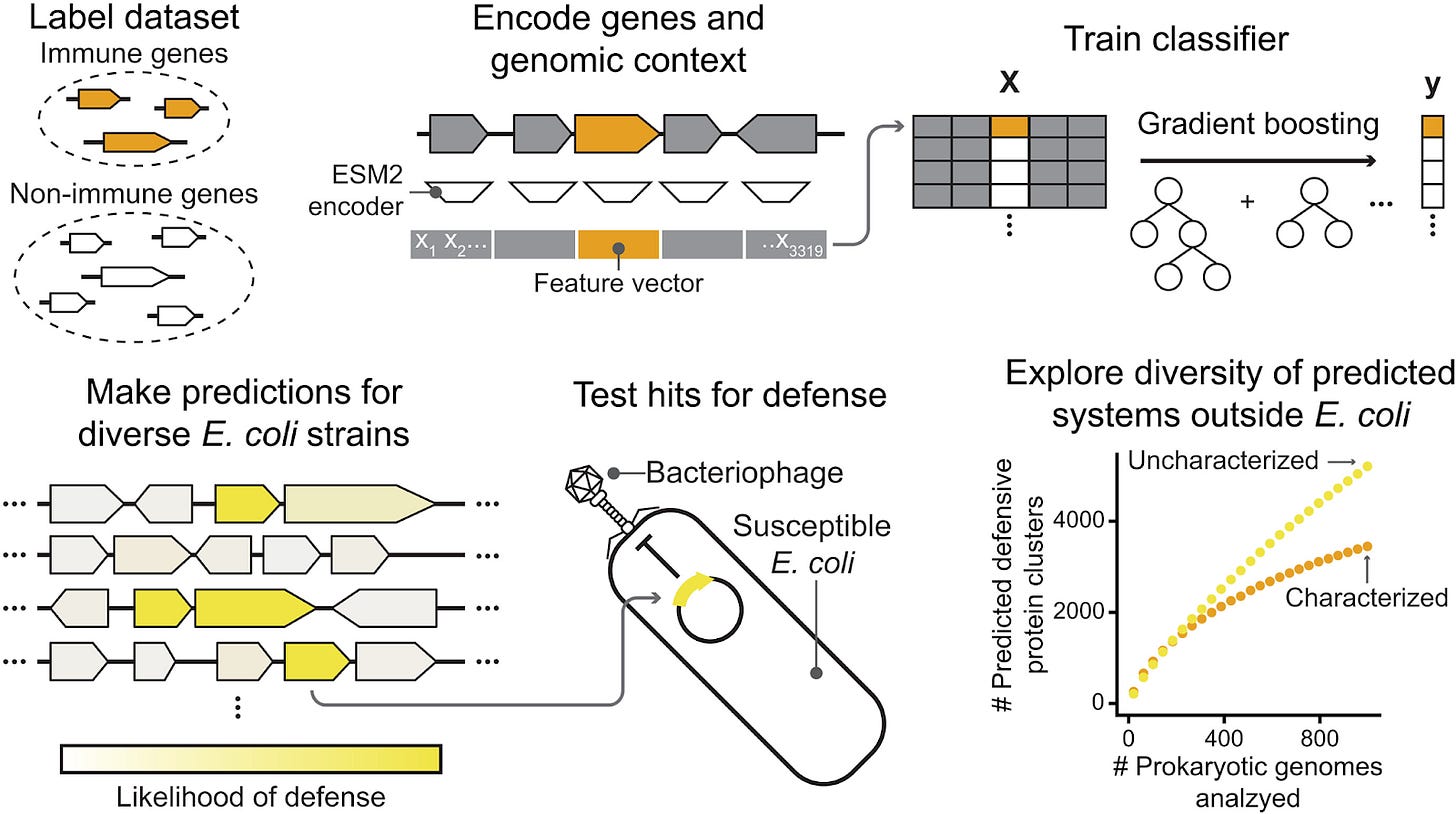
To train DefensePredictor, the authors first needed to collect a dataset of bacterial defense proteins (positive targets) and non-defense proteins (negative targets) to train their classifier. They utilized DefenseFinder, a homology-based method for discovering bacterial defense proteins, across 17,000 genomes to identify 244,000 proteins with putative defense function. For the non-defense proteins, they searched for proteins with gene ontology terms with non-defense function, such as translation or membrane transport, ending up with 14 million putative non-defense genes. After clustering by 30% sequence identity and 80% reciprocal coverage, they finished this step with 15,000 defense proteins and 186,000 non-defense proteins.
To classify whether a protein is a putative bacterial defense gene, DefensePredictor utilizes the ESM2 embedding of the gene itself, the two genes upstream, and the two genes downstream. The authors average across the whole 640-dimensional embedding that ESM outputs for each residue to get a 640-dimensional embedding representing that protein. They concatenate this embedding for the 5 genes into a 3200-dimensional vector, along with 119 computed genomic features, to construct the input to a gradient boosted classifier.
They constructed a stringent cross-validation scheme to ensure that the model isn’t just searching for homologous sequences. They split the dataset into 5 folds, performing an all-by-all MMseqs2 profile search to cluster all of the proteins in the dataset. Each set of homologous proteins was assigned to a single fold - so the model was never tested on a protein family it had already seen during training. On a 5-fold cross-validation with this dataset, their classifier achieved an average precision (AP) of 0.86, outperforming a guilt-by-association method (AP = 0.63) and method utilizing cosine similarity of ESM2 embeddings alone (AP = 0.51). In fact, DefensePredictor can be seen as the combination of the two methods. Just like the guilt-by-association method, DefensePredictor derives much of its power by identifying defense and defense-related proteins in the 4 neighbors around the target gene of interest. Secondly, Inspection of the model using SHAP values revealed that the most important feature of DefensePredictor was the ESM2 embedding of the target gene itself. Because DefensePredictor uses a gradient boosting model to parse ESM2 embeddings, the lower AP of the cosine similarity baseline likely reflects the limitations of that specific geometric comparison rather than a lack of signal within the embeddings themselves.
Most importantly, the authors applied DefensePredictor to search through 69 E. coli genomes in search of new anti-phage defense proteins and experimentally characterized them. They identified 512 protein clusters that were not captured by DefenseFinder. Of those 512 clusters, 103 piqued their interest: they had no detectable homology or were adjacent to known defense proteins. The model effectively detected ‘orphaned’ defense proteins. This group is notably enriched with rare sequences found in very few E. coli genomes, contrasting with the more common distribution of established defense proteins. They experimentally tested 94 of these proteins in E. coli MG1655 and challenged the strain with a phage infection. 42/94 of these proteins successfully helped the strain fight off phage infection! Further characterization of these validated systems revealed a diverse array of protein families and biochemical functions, including metallophosphatases, HipA-family kinases, and PIN ribonucleases, some of which share striking structural homology with eukaryotic innate immune components. Finally, they applied DefensePredictor to 1,000 diverse prokaryotic genomes to uncover over 5,000 uncharacterized defense protein clusters, providing a rich new repertoire of candidates for the development of next-generation genome editing and biotechnological tools.
Temporal AI model predicts drivers of cell state trajectories across human aging [Ortega et al., bioRxiv, April 2026]
Why it matters: MaxToki is a “temporal AI” model capable of reasoning across cell state trajectories to predict interventions that can induce cell state transitions. The authors specifically adapt the model to predict drivers of rejuvenating or pro-aging processes and validate top predictions in mice models.
The past few years have been witness to a steady stream of foundation models like Geneformer, scGPT, and STATE to name a few. One crucial limitation of these models when trying to study gene networks and cellular dynamics is that they can only reason over one cell state at a time when considering a perturbation or change. However, such approaches are not suitable to modeling long timescale processes like development, disease, and aging. In this work, Ortega et al. detail the development of MaxToki, a temporal model trained capable of predicting interventions to induce desired cell state transitions. Specifically, MaxToki was able to infer age acceleration in a variety of unseen disease contexts and predict age-promoting or rejuvenating perturbations that were corroborated by experimental validation.
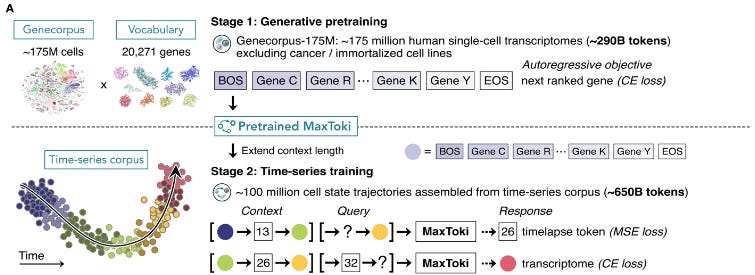
MaxToki was trained over two stages, with the first stage consisting of generative pretraining. This initial training utilized Genecorpus-175M, an assembled dataset of approximately 175 million single-cell transcriptomes spanning various healthy and diseased tissue states. Notably, the team excluded malignant and immortalized cell lines to prevent the model learning from mutated, gain-of-function behaviors. This data was fed to the model in the form of rank value encodings, where genes were ranked based on relative expression within a cell and then scaled across the whole corpus to prioritize highly dynamic genes like transcription factors over others like housekeeping genes. MaxToki itself used an autoregressive transformer decoder architecture and was tasked with predicting the next gene within an input cell during pretraining. The second stage of training was designed to give the model a sense of time by massively expanding the context length to allow the model to process sequences of multiple cells. Specifically, two different prompting tasks were used - could the model predict the time between a “context” and future “query” cell state or predict a future cell state given a context and some queried time lapse.
To gear MaxToki towards aging, the team assembled Genecorpus-Aging-22M. Since tracking the same cells across a human lifespan is not feasible, the authors compiled nearly 22 million cells across 600 cell types from nearly four thousand donors made up of newborns to people over ninety. Using this data, they were able to simulate nearly 100 million aging trajectories and held out certain cell types for downstream testing. Initial evaluations showed that MaxToki was able to successfully predict the time between cells even with scrambled orders, with strong performance across unseen patient ages and cell types. When tasked with generating cell states for held-out age ranges, the model was able to generate transcriptomes whose nearest ground-truth neighbors matched the queried age. Ablation studies showed that the model heavily relied on the rank order of genes and relative expression differences to infer cell states and trajectories. Interestingly, the model’s attention heads were found to pay significant attention to transcription factors, showing that the model could learn the main drivers of cell state transitions.
Finally, the authors used MaxToki to predict if the (computational) deletion of genes would result in a cell state being younger (rejuvenating) or older (pro-aging). The model correctly predicted that repressing GSN had a rejuvenating effect in cardiac fibroblasts; similarly, predictions that the repression of ZBTB16 in capillary endothelial cells was pro-aging was confirmed by wet lab results where perturbed cells showed greater senescence. In cardiomyocytes, MaxToki predicted several pro-aging targets that, when overexpressed experimentally in lab-grown human cells, resulted in major functional defects, delayed calcium cycle kinetics, and activated inflammatory pathways. To demonstrate that these results could be validated in a full living organism, the authors used viral vectors to deliver top predicted pro-aging genes into the hearts of adult mice which showed significantly worse cardiovascular function within six weeks. The mix of computational and experimental results prove that MaxToki’s temporal reasoning framework is capable of generalizing to unseen cellular trajectories and can be successfully adapted to studying processes like aging and disease. The authors point to capturing longitudinal disease data as their next step to see if temporal models can understand intersecting disease axes.
Programmable macromolecule delivery via engineered trogocytosis [Chen et al., Nature Cell Biology, April 2026]
Why it matters: Delivering macromolecules into specific cell types remains a central bottleneck across gene editing, cell therapy, and protein therapeutics. Existing approaches, such as viral vectors, liquid nanoparticles, or extracellular vesicles, offer scalability yet limited control over cell-type specificity and intracellular fate. Cell-cell transfer mechanisms such as trogocytosis provide a direct, contact-dependent alternative, but have been constrained by poor cargo control, limited functional delivery, and unclear generalizability.
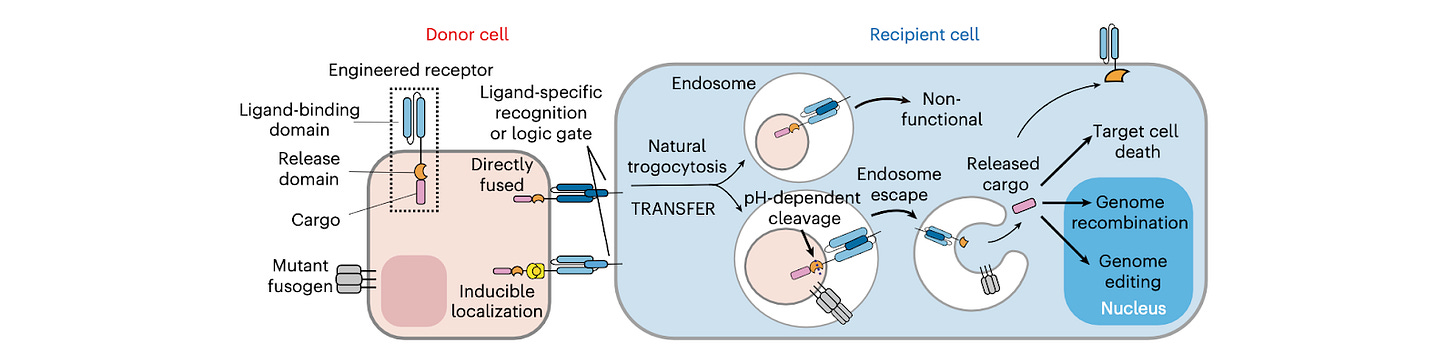
This paper establishes engineered trogocytosis as a programmable delivery system. Chen et al. design donor cells expressing synthetic receptors that bind defined ligands on recipient cells, enabling targeted transfer of membrane-associated proteins during cell-cell contact. They identify key principles that enable efficient transfer and functional integration of cargos, such as receptor affinity, inducible cargo localization, and pH-responsive membrane fusion.
Mechanistically, transferred cargos enter recipient cells through an endosome-dependent pathway, where acidification enables membrane fusion and release of functional payloads. Imaging and compartment analysis show that cargo functionalization occurs early in the endosomal pathway, with no bias toward degradation versus recycling routes, allowing a substantial fraction of transferred material to remain functional.
The system is generalized into TRANSFER, a modular platform that supports programmable targeting and payload delivery. TRANSFER can integrate multiple ligand inputs for conditional targeting, deliver large protein cargos, and enable functional outputs including genome editing in recipient cells. The approach is compatible across multiple cell types and operates through conserved cellular trafficking mechanisms.
Overall, this work reframes trogocytosis into a designable cell-cell delivery interface. By coupling specificity at the cell-contact level with intracellular release control, it expands the design space for targeted biologic delivery and suggests a path toward programmable, cell-mediated therapeutic systems.
Notable deals
Anthropic acquires stealth startup Coefficient Bio in $400M stock deal to build out presence in the healthcare space. The Dimension-backed startup was reportedly pursuing development of “artificial superintelligence for science”, as reported by Eric Newcomer in his post, one of the original sources of coverage of the still highly secretive deal. The acquisition follows the recent launch of Claude Life Sciences, wherein Anthropic is seeking to model the drug development process from end to end, spanning discovery all the way through clinical trials. Coefficient, on the other hand, was launched a mere eight months ago (~2 months prior to Claude’s life sciences debut) and, at the time of the deal announcement, consists of a team of less than ten individuals, all of whom will purportedly be joining Anthropic’s Health Care Life Sciences team. This deal marks AI giants as a potential new class of biotech acquirer, signalling a possible expansion of exit opportunities—especially for tech-forward life science companies—beyond traditional pharma buyers.
Sidewinder Therapeutics announces the close of oversubscribed $137M Series B co-led by Frazier Life Sciences and Novartis Venture Fund. The funding will be used to progress the company’s next-generation bispecific antibody drug conjugates (ADCs) into the clinic for solid state tumors in a variety of oncology indications which currently lack significant existing treatment options but affect substantial patient populations. Potential applications include lung, head, and neck squamous cell carcinomas, and GI cancers such as colorectal cancer (which has notably been on the rise in younger populations in recent years). Sidewinder emphasizes precise targeting of highly-expressed, tumor-specific receptor co-complexes present on solid state tumors as a key part of their approach which allows for greater specificity and differentiation in treatment delivery—targeting cancerous cells while preserving normal cells in proximity. Eric Murphy, Sidewinder’s CEO, professes the belief that ADCs are at a critical inflection point, poised for technological breakthrough—which Sidewinder hopes to usher in as they prepare for their lead program to enter clinical trials in 2027. Other participants in the fundraising include: Orbimed (existing investor), and new investors, Life Sciences at Goldman Sachs Alternatives, DCVC Bio, Samsara BioCapital, Longwood Fund, Astellas Venture Management and Alexandria Venture Investments.
Life Biosciences raises $80M Series D to support cellular rejuvenation pipeline as lead asset just commences Phase 1 trials. Life’s lead candidate, ER-001, is currently being assessed for safety and tolerability in the clinic in patients with optic neuropathies, specifically open angle glaucoma (OAG) and non-arteritic anterior ischemic optic neuropathy (NAION). Such indications present with damage to retinal ganglion cells (RGCs) which causes permanent vision impairment due to the lack of capacity for RGCs to regenerate. Optic neuropathies as they stand today have substantial unmet need with respect to current treatments which fail to offer vision preservation or restoration, much like many other chronic, aging-related diseases. As such, the company’s PER platform seeks to address this gap for optic neuropathies and other aging indications via partial epigenetic reprogramming using several transcription factors: OCT4, SOX2, and KLF4 (OSK). Investors in the round are presently undisclosed.
Stipple Bio emerges from stealth with $100M oversubscribed Series A co-led by RA Capital, a16z Bio+Health, and Nextech Invest. Stipple’s name originates from the company’s precision approach to oncology of mapping thousands of subtle molecular points to reveal tumor-specific epitopes ripe for therapeutic targeting via their proprietary Pointillist Platform. Funding from the round will serve to advance Stipple’s lead candidate, STP-100, a novel ADC with “a clinically validated linker payload incorporating tumor-specific binders” into multiple clinical trials; it will also support operational runway through 2029. A major goal of the company’s next-generation precision ADCs is sparing normal cells in the targeted tumor vicinity, similar to aims professed by Sidewinder Therapeutics which also raised this week.
Neurocrine Biosciences to acquire Soleno Therapeutics in an equity transaction totaling $2.9B. In completing this deal, Neurocrine is seeking to expand and diversify their endocrinology and rare disease portfolio with the addition of Soleno’s VYKAT XR (diazoxide choline), a first-in-class and only existing therapeutic for hyperphagia, a persistent hunger resulting from abnormalities in chromosome 15, resulting in compulsive food-seeking and the defining characteristic of Prader-Willi Syndrome (PWS). Launched in Q2 2025, VYKAT has experienced strong early adoption resulting in significant returns for Soleno, with Neurocrine’s press release reporting $190M in revenue for 2025 ($92M just from Q4). Neurocrine projects long-term returns from the drug with potential for market expansion with broader utilization as well as delivering life-changing treatment outcomes for impacted patients. VYKAT now marks Neurocrine’s third marketed, first-in-class drug alongside INGREZZA, a vesicular monoamine transmitter 2 treatment for tardive dyskinesia and Huntington’s-associated chorea, and CRENESSITY, a therapeutic targeting classic congenital adrenal hyperplasia resulting from 21-hydroxylase deficiency.
China-based Syneron Bio raises $150M Series B to further macrocyclic peptide development platform.The round was led by an undisclosed international life sciences fund along with Decheng Capital and CDH GCV, and follows a nearly $100M Series A just a few months earlier as well as a multibillion-dollar biobucks deal with AstraZeneca penned one year ago. Macrocyclic peptides are currently generating significant attention from pharma as evidenced by another $1.7B deal between Novartis and US-based Unnatural Products just this past February. The modality is being referred to as a potential “goldilocks” drug class due to their capability to target historically difficult-to-treat disease pathways eluding small molecules and biologics, according to Fierce Biotech coverage of the fundraise. Disclosed use of funds from the round is somewhat vague, being reportedly attributed as going toward advancing Syneron’s oncology, autoimmune, metabolic, and rare disease programs. Other new investors in the round include: a wholly owned subsidiary of the Abu Dhabi Investment Authority, True Light Capital, Qiming Venture Partners, BioTrack Capital; alongside existing investors: AstraZeneca, LAV, Sinovation Capital, 5Y Capital, GL Ventures, Biotech Development Fund and Lenovo Capital.
In case you missed it
What we listened to
What we liked on socials channels
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @decodingbio.