BioByte 151: Protein-Complexa Tackles Fully Atomistic Binder Design, CAR-As Help Clear Amyloid-β Plaques, How Granule Cells Enable Effective Learning, and the Tradeoff Between Regeneration and Cancer
Welcome to Decoding Bio’s BioByte: each week our writing collective highlight notable news—from the latest scientific papers to the latest funding rounds—and everything in between. All in one place.
What we read
Blogs
The Regeneration-Cancer Tradeoff [Karlin Compton, Proof Of Compton, March 2026]
Humans are deficient in true regenerative capabilities, being limited to repairing blood, gut lining, skin, and parts of the liver rather than the entire body. The regeneration that does occur mostly proceeds through signaling via growth factors such as FGF and VEGF, which can help to temporarily induce a state closer to that of stem cells near damaged tissue until healing is complete. In contrast, these same growth factors and signaling modes often remain uninhibited or permanently active in cancers, netting them the description of “the wound that never heals”. Given these observations, Compton argues the limitation on humanity’s ability to truly regenerate is due to the similar biological machinery and pathways shared between regenerative capabilities and the growth and propagation of cancers. Relative to species which can regenerate more fully, human evolution proceeded along a more conservative path.
The epithelial-mesenchymal transition (EMT) and dedifferentiation are two examples Compton provides for the similarity between regeneration and cancer. In the EMT, the adherence of some cells to the epithelial layers is relaxed, affording them migratory capabilities. This process is an essential part of transporting necessary cells to the site of a wound, but it is also one of the ways in which cancers are able to metastasize, making too much promiscuity in the EMT process a significant danger. In dedifferentiation, specialized cells are transformed into more labile forms, allowing them to participate in regeneration. However, the modifications that occur during this process closely resemble that of cancer cells such as epigenetic remodeling and a loosening of the cell’s regulatory architecture. Labile cells which can readily propagate could either be a recipe for limb regeneration or cancer.
Compton suggests that in regenerative creatures (like zebrafish, salamanders, and planaria) processes like EMT and dedifferentiation are highly regulated such that when they have fulfilled their purpose, they are shut down, drawing the conclusion that the difference between regeneration and cancer arises from differences in control and regulation. It will be interesting to follow along as more research is done to understand what controls these regenerative behaviors, how it relates to cancer formation, and what specific mechanisms are at play which inhibit cancer formation and propagation.
Papers
Scaling Atomistic Protein Binder Design with Generative Pretraining and Test-Time Compute [Didi et al., preprint, March 2026]
Why it matters: Proteina-Complexa is a fully atomistic binder design tool that marries conditional generation methods and structure-based hallucination tools to deliver significant performance gains across a range of binder design challenges. The framework is also accompanied with rigorous wet lab validation that showcases its potential beyond in silico benchmarking.
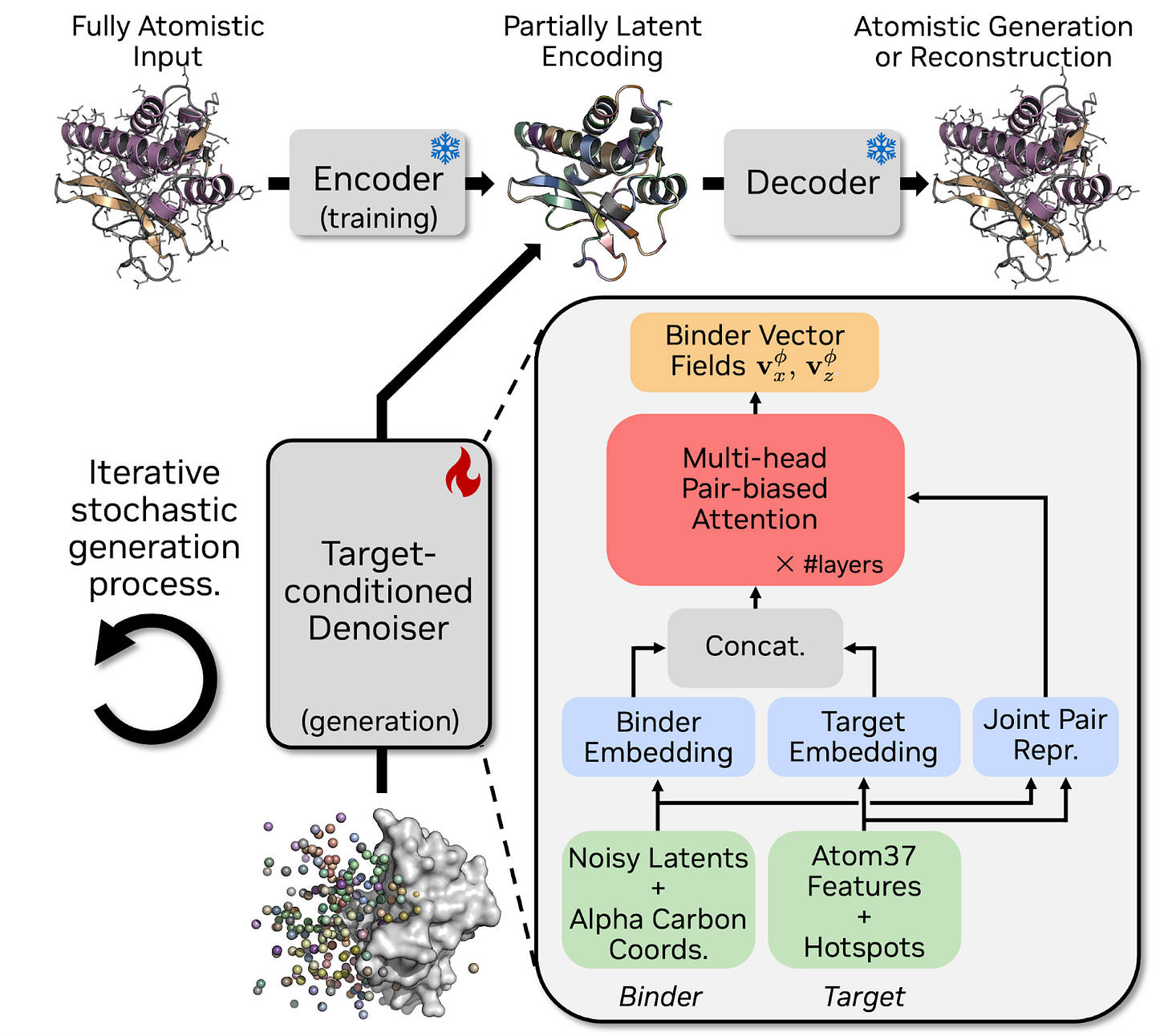
Building on their previous work from La Proteina, the Fundamental Generative AI Research (GenAIR) Group at NVIDIA released Proteina-Complexa, a new framework to tackle fully atomistic binder design challenges. The development of Proteina-Complexa is grounded in the idea that the current dichotomy of binder generation methods is ripe for optimization. Briefly, binder generation methods fall under two classes - generative or hallucination-based - with the former (like RFDiffusion) approach relying on conditional generation processes while the latter (like BindCraft) uses gradients from structure predictors to iteratively refine a candidate design. The authors argue that this division is in stark contrast to the integrated framework of generative priors and inference-time scaling of leading language and vision models. In this paper Didi et al. present Proteina-Complexa as “the first binder design framework that unifies a strong flow-based base generative model with flexible inference-time optimization utilizing the generative prior, combining the strengths of both.”
Proteina-Complexa reframes La-Proteina’s partially latent flow matching objective and applies it to conditional binder design. Specifically, the model uses an autoencoder to encode alpha-carbon backbone coordinates to control basic structure and uses latent variables for each residue to control amino acid properties and side chain geometries. To adapt this framework for binder design, the team introduced a target-conditioned denoiser that concatenates embedded target features with the binder’s latent representation before passing them to transformer layers that can process the joint embeddings. To leverage the benefits of structure-based hallucination methods, Proteina-Complexa uses structure prediction confidence metrics and interface bond energies to steer the generative diffusion process. In parallel, the team also used a range of test-time compute scaling approaches like growing the candidate pool, balancing exploration and exploitation with Monte Carlo tree searches, and varying reward-based strategies for goals like increasing interface hydrogen bonds.
Apart from model architecture developments, other significant contributions include the development of the Teddymer Dataset, meant to bridge the gap in experimentally resolved multimers for paired binder-target training data. The team split the AlphaFold Database (AFDB) into individual domains and curated a set of synthetic dimers, yielding a dataset (Teddymer) much larger than the standard PDB. Proteina-Complexa was trained across four stages of data, starting with AFDB monomers followed by Teddymer dimers, before moving on to PDB multimers and protein-ligand pairs from the PLINDER dataset to give the model access to a wide variety of binder-target pairs. The benefits of this training regime were evident from both in silico benchmarks and an extensive wet lab validation campaign. Across a range of targets, the model generally outperformed conditional generation methods like RFDiffusion and Protpardelle-1c, while also outperforming tools like BoltzDesign, BindCraft, and AlphaDesign across a range of inference-time scaling approaches. Finally, Proteina-Complexa’s predictions showed promising wet lab performance, with the model demonstrating high hit rates and picomolar affinities for certain targets, while also designing the first known successful de novo carbohydrate binders. The wet lab performance is outlined in a separate manuscript. In summary, the combination of promising performance across computational and wet lab benchmarks reaffirms the potential of generative design methodologies that can merge conditional generation and hallucination-based approaches.
Targeting amyloid-β pathology by chimeric antigen receptor astrocyte (CAR-A) therapy [Chen et al., Science, March 2026]
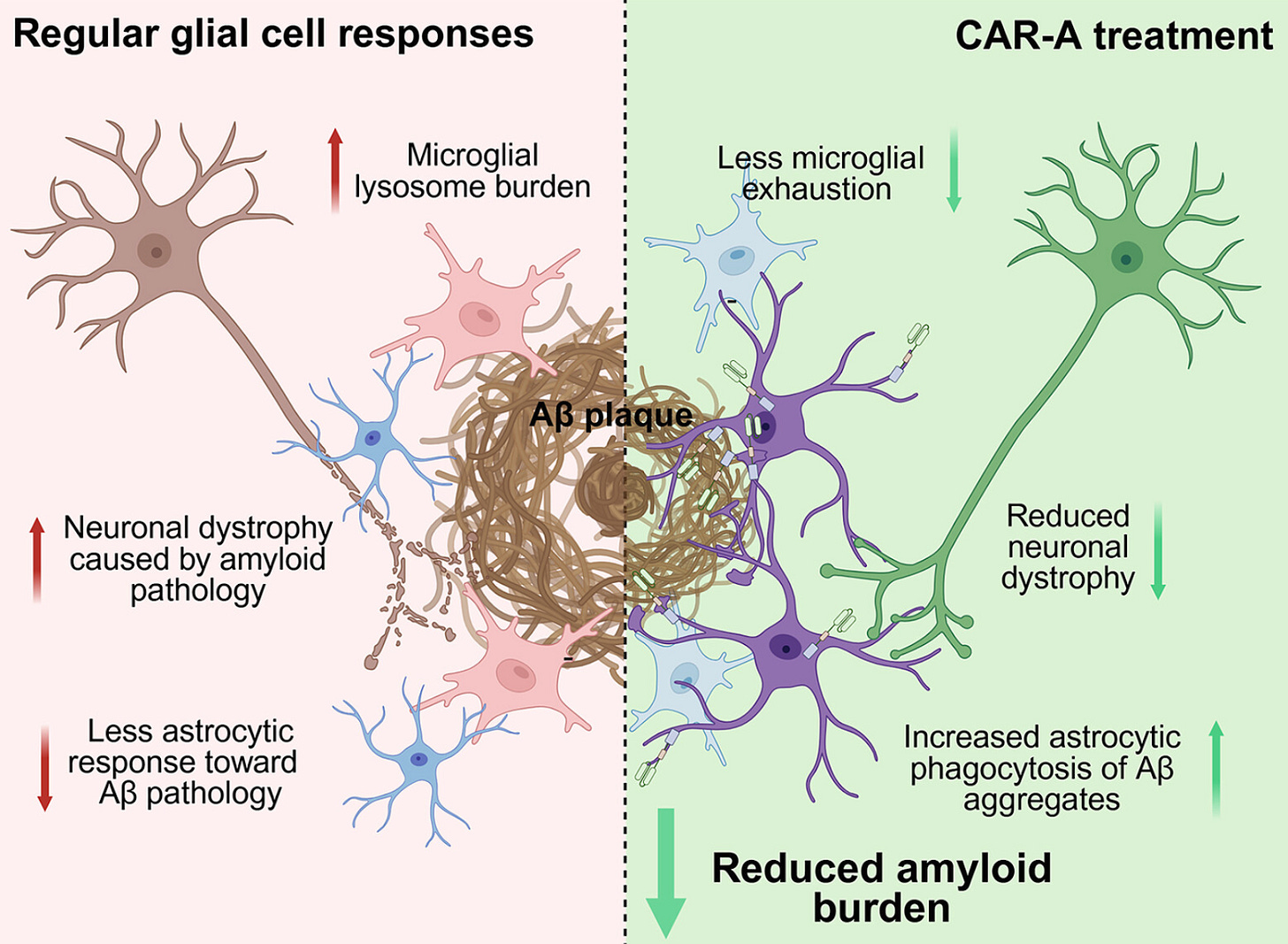
Why it matters: Chen et al. develop a new strategy to clear amyloid-β plaques, opening up a new avenue for anti-Aβ therapeutics. By arming astrocytes with chimeric antigen receptors (CARs), they enable these cells to degrade Aβ plaques through phagocytosis, leading to reduced plaque burden in mouse models.
Even though anti-amyloid antibodies have now reached the clinic for Alzheimer’s treatment, it remains unclear how much plaque removal translates into meaningful disease modification. These therapies also come with real tradeoffs, including repeated high-dose administration, narrow treatment windows, and safety concerns such as amyloid-related imaging abnormalities. Instead of relying on antibodies to recruit endogenous phagocytes, the authors asked whether they could directly reprogram brain glia to recognize and clear amyloid-β themselves. Others have explored similar CAR-based approaches in microglia or myeloid cells, but these strategies face a key challenge: resident microglia are difficult to replace at scale, and engineered myeloid cells have not demonstrated durable, brain-wide activity in vivo. The authors instead turned to astrocytes, which are broadly distributed throughout the brain, already participate in phagocytic processes, and are accessible via CNS-tropic AAV delivery.
To build the system, the team fused anti-amyloid single-chain antibody fragments to intracellular signaling domains from phagocytic receptors, creating four FcRγ-independent CAR designs for astrocytes. In their initial screen, they paired a crenezumab-derived scFv (targeting Aβ) with signaling domains from MEGF10, Dectin1, MERTK, or CD3ζ. After validating function in vitro, they moved in vivo to test whether these engineered astrocytes could meaningfully reduce amyloid burden and associated pathology. A single AAV-delivered dose administered after plaque formation reduced amyloid burden by about half in 5xFAD mice and decreased neuritic dystrophy, while earlier administration, prior to plaque deposition, strongly suppressed subsequent amyloid accumulation and related pathology.
While this in vivo engineering strategy effectively reduces plaque burden, the authors highlight several limitations and areas for further development. Despite robust amyloid clearance, synaptic rescue was only partial, and the therapy may lead to overpruning of healthy synapses. Improvements in overall neural health were also limited, though this remains difficult to interpret given known constraints of the mouse model and broader uncertainty in the field around whether amyloid clearance alone is sufficient to restore function. Still, as a first demonstration of CAR-armed astrocytes operating in vivo, this represents a compelling proof of concept, and it will be exciting to see how the field iterates on construct design, safety, and cell-type targeting moving forward.
Granule cells reorient cortical manifolds to separate contexts but preserve their geometry [Garcia-Garcia et al., bioRxiv, March 2026]
Why it matters: To learn effectively, animals must generalize across related experiences while still distinguishing between them – a fundamental computational tradeoff. Generalization is supported by low-dimensional neural manifolds found throughout the neocortex which accelerate learning by constraining activity to a small number of task-relevant axes. Conversely, context separation is classically attributed to neural expansion layers – most famously, cerebellar granule cells (GrCs) – that project overlapping inputs into high-rank feature spaces to separate them. However, high-rank expansion ‘shatters’ the smooth structure required for continuous prediction and generalization, creating an apparent paradox. Garcia-Garcia et al. resolve it by simultaneously imaging premotor cortical output neurons (layer 5 pyramidal tract cells, L5PT) and GrCs during parallel skill learning, revealing that GrCs reject high-dimensional expansion. Instead, they perform structured affine transformations – rotating cortical manifolds apart across contexts while rigorously preserving their low-rank geometry within each.
Using a custom dual-site two-photon microscope, the authors tracked the same L5PT and GrC populations across two tasks with shared temporal structure (action → delay → reward) but distinct sensorimotor demands (forelimb reach vs VR locomotion) in 9 mice. Within each task, GrCs faithfully preserved cortical manifold structure: comparably dense, temporally organized, and low-rank (~4-5 effective dimensions by participation ratio, a measure of dimensionality based on eigenvalue spectra). Across tasks, however, the populations diverged. L5PT modes generalized strongly (cross-task r = 0.83), while GrC modes remapped nearly completely (r = 0.24) – despite stable coupling between the two regions. Representational similarity analysis of neural activity showed GrCs preserved within-task cortical geometry with high fidelity (Spearman ρ = 0.82) but diverged across tasks (ρ = 0.48). GrC manifolds rotated significantly further apart than L5PT manifolds across contexts, and this separation grew with dual-task proficiency (ρ = 0.6): as behavior converged across tasks, GrC representations became more distinct.
To test whether this strategy confers a computational advantage, the authors simulated alternating two-task learning in a cerebellar-like circuit, comparing three GrC architectures: one that simply relays cortical geometry, one that shatters it via classical sparse expansion, and one that rotates manifolds apart (matching their data). The relay learned quickly but suffered catastrophic forgetting between tasks due to overlapping representations. Sparse expansion avoided forgetting but learned very slowly – shattering the geometry destroyed the smooth structure needed for temporal prediction. The rotation model achieved both: fast learning and resistance to interference. The authors propose that context-dependent rescaling of inputs to GrCs could generate these rotations, but this mechanism remains unvalidated. Interesting open questions follow from these experiments – including whether the motif extends to other pathways and expansion architectures (ex. piriform cortex, dentate gyrus) and quantifying the manifold packing capacities for different contexts. If this generalizes, coherent manifold reorientation may represent a broadly applicable blueprint for continuous prediction and control for neuroscientific and engineered systems.
Notable deals
Unnatural Products raises a $45M Series B round led by The Venture Collective (TVC). The company is seeking to unlock an entirely new class of drugs: oral synthetic macrocyclic peptides. Previously only able to be targeted via costly injectables, Unnatural Products is using their proprietary platform to go after historically undruggable targets, having inked an impressive partnership with Novartis worth $100M upfront with $1.7B in potential milestones to go after such elusive targets in the cardiovascular domain just last month. The company has additional established collaborations with Merck, BridgeBio, and argenyx spanning multiple therapeutic areas whilst advancing their own pipeline of treatments for cardiometabolic, autoimmune, and inflammatory indications. Other investors participating in the round include argenx, Droia Ventures and existing investors Merck Global Health Innovation Fund, Artis Ventures, and First Spark Ventures.
Vima Therapeutics announces $40M Series A extension as first patient is dosed in Phase 2 dystonia study. The movement disorders-focused biotech is aiming at a potential first-in-class therapeutic for dystonia and Parkinson’s disease, which share underlying disease biology consisting of dopaminergic and acetylcholinergic signalling imbalances in the brain. Vima’s lead candidate, VIM0423, is a once-daily oral drug intended to selectively target muscarinic cholinergic receptors in the brain, a target established as effective, but one that has been limited by tolerability issues. VIM0423 seeks to address these tolerability challenges whilst still maintaining maximum efficacy. The potential treatment was awarded Fast Track designation by the FDA and is moving into Phase 2 clinical trials, having just completed a two-part Phase 1 study assessing pharmacokinetics, safety, tolerability, and dose titration. Funding from this round—which saw participation from new investor, Frazier Life Sciences, as well as from existing investors, Atlas Ventures, Access Industries, and Canaan Partners—will support this ongoing Phase 2.
Crossbow Therapeutics closes $77M Series B led by Taiho Ventures and Arkin Bio Capital. Armed with a portfolio of novel precision antibody drugs, Crossbow is aiming at treating a broad range of cancers. Their proprietary immunotherapies—dubbed T-Bolt™ molecules—consist of T cell engagers (TCE) targeting peptide human leukocyte antigen (pHLA) located on cancer cells via T cell mimetics (antibodies that resemble T cell receptors). This funding will bolster the company’s lead candidate, CBX-250, a first-in-class TCE through the CROSSCHECK-001 Phase 1 clinical trial which is set to evaluate the therapeutic in patients experiencing relapsed and refractory myeloid malignancies across acute myeloid leukemia, chronic myeloid leukemia, myelodysplastic syndromes, and chronic myelomonocytic leukemia. The Series B will additionally serve to advance other promising TCE assets such as CBX-663 which targets a telomerase reverse transcriptase (TERT)-derived pHLA for the treatment of multiple hematologic and solid tumors. CBX-663 is currently slated for IND submission. Other investors in the round include new adds: Sixty Degree Capital, Hamilton Square Partners Management LP, LifeLink Ventures, Libbs Ventures, and Blood Cancer United’s Therapy Acceleration Program (TAP), as well as existing investors MPM BioImpact, Pfizer Ventures, BVF Partners, Polaris Partners, Eli Lilly and Company, and Mirae Asset Venture Investment.
With a $77.5 million, oversubscribed Series A, R1 Therapeutics launches to treat chronic kidney disease. In addition to the announcement of the raise, R1 also publicized the acquisition of the development and commercialization rights to AP306—a pan phosphate transporter inhibitor—outside of China from Alebund Pharmaceuticals. The asset is scheduled to enter Phase 2b trials for hyperphosphatemia in patients with chronic kidney disease later in 2026. Phase 2a results showed significant reduction in serum phosphate levels thanks to the ability of AP306 to block the active transport of phosphate, unlike other therapies which only block passive transport. Abingworth, DaVita Venture Group, and F-Prime led the round while Curie.Bio, SymBiosis, and U.S. Renal Care also participated.
In case you missed it
Breakout Ventures Launches Fund III with $114M to Continue Building the Future Powered by Science
Droplet Biosciences Accelerates Residual Cancer Detection with NVIDIA
Exploring the Scaling Laws of Consumer EEG: Introducing Alljoined-1.6M
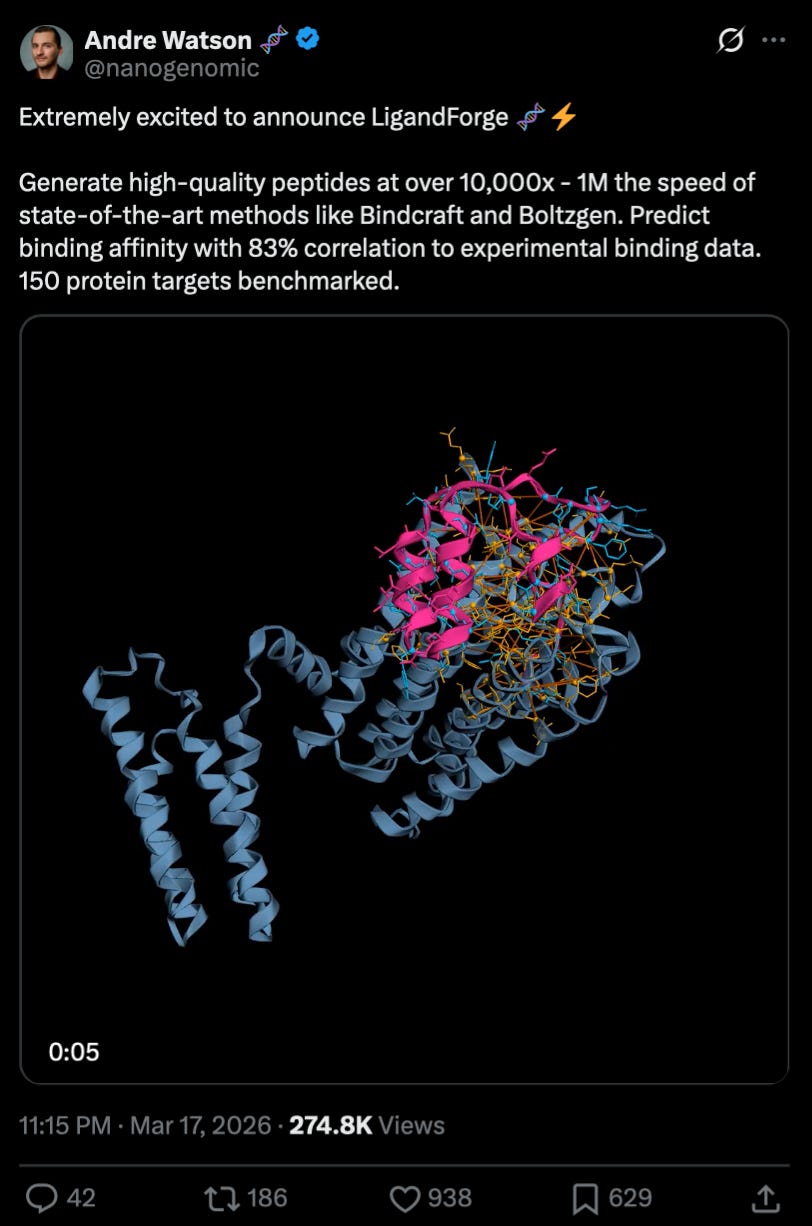
What we liked on socials channels
Bonus content: everyone online is buzzing about the new LinkedIn Speak translator so we had to check it out ourselves (don’t worry we love decoding! ok now back to proofreading 🤓)
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @decodingbio.